Details
Structure visualisation
Entry information
| Complex | |
| AACDB_ID: | 6978 |
| PDBID: | 4NBY |
| Chains: | C_A |
| Organism: | Clostridioides difficile NAP07, Lama glama |
| Method: | XRD |
| Resolution (Å): | 2.08 |
| Reference: | 10.1074/jbc.M113.505917 |
| Antibody | |
| Antibody: | A20.1 VHH |
| Antibody mutation: | No |
| INN (Clinical Trial): | |
| Antigen | |
| Antigen: | Rho-glucosylating toxins TcdA2 |
| Antigen mutation: | No |
| Durg Target: | |
Sequence information
Antibody
Chain: C
Mutation: NULL
| >4NBY_C|Chain B, C|A20.1 VHH|Lama glama (9844) QPAMAQAQVQLVESGGGLAQAGGSLRLSCAASGRTFSMDPMAWFRQPPGKEREFVAAGSSTGRTTYYADSVKGRFTISRDNAKNTVYLQMNSLKPEDTAVYYCAAAPYGANWYRDEYAYWGQGTQVTVSSGQAGQGSEQKLISEEDLNHHHHHH |
Antigen
Chain: A
Mutation: NULL
| >4NBY_A|Chain A|Cell wall-binding repeat protein|Clostridium difficile (525258) MHHHHHHSKAVTGMRTIDGKKYYFNTNTAEAATGWQTIDGKKYYFNTNTSIASTGYTIINDKHFYFNTDGIMQIGVFKGPDGFEYFAPANTDANNIEGQAIRYQNRFLYLHDNIYYFGNNSKAATGWVTIDGRRYYFEPNTAIGANGYKIIDNKNFYFRNGLPQIGVFKGPNGFEYFAPANTDANNIDGQAIRYQNRFLHLLGNIYYFGNNSKAVTGWQTINGNMYYFMPDTAMAAAGGLFEIDGVIYFFGVDGVKAPGIY |
Interaction
1、Solvent accessible surface areas (SASA) were calculated (Naccess V2.1.1) for each residue in antibody and antigen, respectively. The residues with SASA loss in binding of more than 1Å2 were classified as interacting residues.
Interacting residues (ΔSASA based)
| Chain residues position delta_SASA
: residuesposition C: GLN8 VAL9 GLY33 ARG34 THR35 SER37 MET38 ASP39 PRO40 SER59 SER60 THR61 ARG63 THR64 TYR66 ALA106 PRO107 TYR108 GLY109 ALA110 ASN111 TRP112 ARG114 ALA118 TYR119 A: ILE74 TYR85 PRO88 ALA89 ASN90 THR91 ASN94 TYR103 ARG106 PHE107 LEU108 TYR109 LEU110 HIS111 ASP112 ASP131 ARG133 TYR135 ARG159 ASN160 |
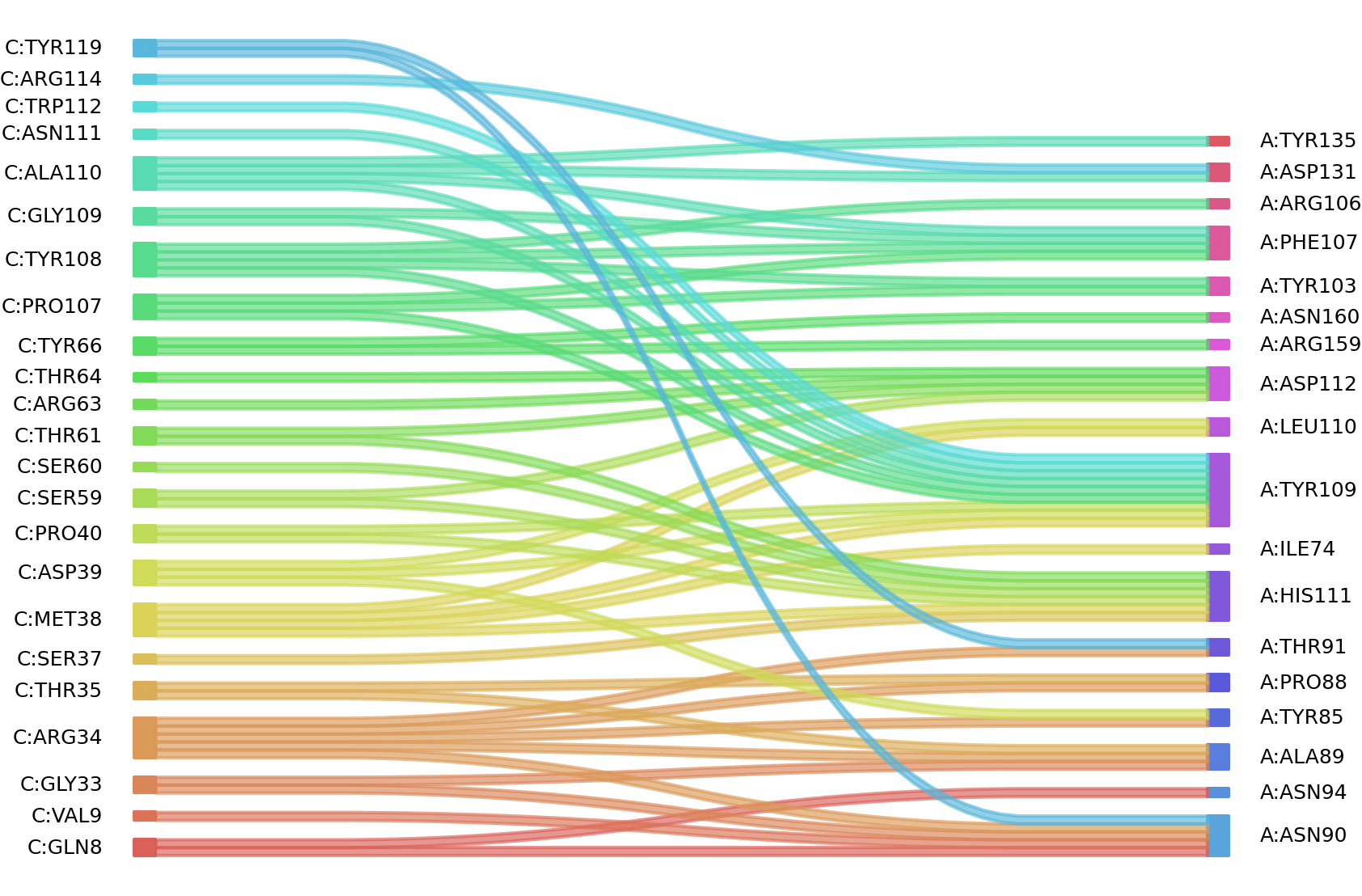
2、We defined interacting paratope-epitope residues by a distance cutoff of < 6 Å . Two amino acids are considered as interacting residues if they have at least one atom within a distance of 6 Å from any atom.
Interacting residues (Atom distance based)