Details
Structure visualisation
Entry information
| Complex | |
| AACDB_ID: | 5652 |
| PDBID: | 7Y6K |
| Chains: | E_B |
| Organism: | Severe acute respiratory syndrome coronavirus 2, Homo sapiens |
| Method: | EM |
| Resolution (Å): | 3.34 |
| Reference: | Not available or To be published |
| Antibody | |
| Antibody: | K202.B scFv |
| Antibody mutation: | No |
| INN (Clinical Trial): | |
| Antigen | |
| Antigen: | SARS-CoV-2 spike glycoprotein RBD |
| Antigen mutation: | Yes |
| Durg Target: | P0DTC2; P0DTC2; |
Sequence information
Antibody
Chain: E
Mutation: NULL
| >7Y6K_E|Chain B[auth E], C[auth I]|scFv region of light chain from K202.B, bispecific antibody|Homo sapiens (9606) EVQVLESGGGLVQPGGSLRLSCAASGFTFSSYDMSWVRQAPGKGLEWVSGISYSGGSTYYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCARDVNFVPLVTFDYWGQGTLVTVSSGGGGSGGGGSGGGGSQSVLTQPPSASGTPGQRVTLSCTGSSSNIGSNSVTWYQQLPGTAPKLLIYDDNQRPSGVPDRFSGSKSGTSASLAISGLRSEDEADYYCGTWDYSLSAYVFGGGTKLTVLGGGGSGGGGSGGGGSQSVLTQPPSASGTPGQRVTISCSGSSSNIGSNAVSWYQQLPGTAPKLLIYADNKRPSGVPDRFSGSKSGTSASLAISGLRSEDEADYYCGAWDSSLNAYVFGGGTKLTVLRTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC |
Antigen
Chain: B
Mutation: R682G/R683S/R685S/F817P/A892P/A899P/A942P/K986P/V987P
| >7Y6K_B|Chain A[auth B]|Spike glycoprotein|Severe acute respiratory syndrome coronavirus 2 (2697049) MFVFLVLLPLVSSQCVNLTTRTQLPPAYTNSFTRGVYYPDKVFRSSVLHSTQDLFLPFFSNVTWFHAIHVSGTNGTKRFDNPVLPFNDGVYFASTEKSNIIRGWIFGTTLDSKTQSLLIVNNATNVVIKVCEFQFCNDPFLGVYYHKNNKSWMESEFRVYSSANNCTFEYVSQPFLMDLEGKQGNFKNLREFVFKNIDGYFKIYSKHTPINLVRDLPQGFSALEPLVDLPIGINITRFQTLLALHRSYLTPGDSSSGWTAGAAAYYVGYLQPRTFLLKYNENGTITDAVDCALDPLSETKCTLKSFTVEKGIYQTSNFRVQPTESIVRFPNITNLCPFGEVFNATRFASVYAWNRKRISNCVADYSVLYNSASFSTFKCYGVSPTKLNDLCFTNVYADSFVIRGDEVRQIAPGQTGKIADYNYKLPDDFTGCVIAWNSNNLDSKVGGNYNYLYRLFRKSNLKPFERDISTEIYQAGSTPCNGVEGFNCYFPLQSYGFQPTNGVGYQPYRVVVLSFELLHAPATVCGPKKSTNLVKNKCVNFNFNGLTGTGVLTESNKKFLPFQQFGRDIADTTDAVRDPQTLEILDITPCSFGGVSVITPGTNTSNQVAVLYQDVNCTEVPVAIHADQLTPTWRVYSTGSNVFQTRAGCLIGAEHVNNSYECDIPIGAGICASYQTQTNSPGSASSVASQSIIAYTMSLGAENSVAYSNNSIAIPTNFTISVTTEILPVSMTKTSVDCTMYICGDSTECSNLLLQYGSFCTQLNRALTGIAVEQDKNTQEVFAQVKQIYKTPPIKDFGGFNFSQILPDPSKPSKRSPIEDLLFNKVTLADAGFIKQYGDCLGDIAARDLICAQKFNGLTVLPPLLTDEMIAQYTSALLAGTITSGWTFGAGPALQIPFPMQMAYRFNGIGVTQNVLYENQKLIANQFNSAIGKIQDSLSSTPSALGKLQDVVNQNAQALNTLVKQLSSNFGAISSVLNDILSRLDPPEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRASANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQSAPHGVVFLHVTYVPAQEKNFTTAPAICHDGKAHFPREGVFVSNGTHWFVTQRNFYEPQIITTDNTFVSGNCDVVIGIVNNTVYDPLQPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRLNEVAKNLNESLIDLQELGKYEQ |
Interaction
1、Solvent accessible surface areas (SASA) were calculated (Naccess V2.1.1) for each residue in antibody and antigen, respectively. The residues with SASA loss in binding of more than 1Å2 were classified as interacting residues.
Interacting residues (ΔSASA based)
| Chain residues position delta_SASA
: residuesposition E: THR28 SER30 SER31 TYR32 ASP33 TYR53 SER54 SER57 ASN74 SER75 ASP99 VAL100 ASN101 PHE102 VAL103 PRO104 LEU105 GLY166 SER167 ASN168 TYR186 ASP187 ASP188 ASN189 GLN190 LYS203 TYR230 B: TYR369 ASN370 SER371 ALA372 PHE374 SER375 THR376 PHE377 LYS378 CYS379 TYR380 GLY381 VAL382 SER383 PRO384 THR385 GLY404 ASP405 VAL407 ARG408 ASN437 GLY502 VAL503 GLY504 TYR505 TYR508 |
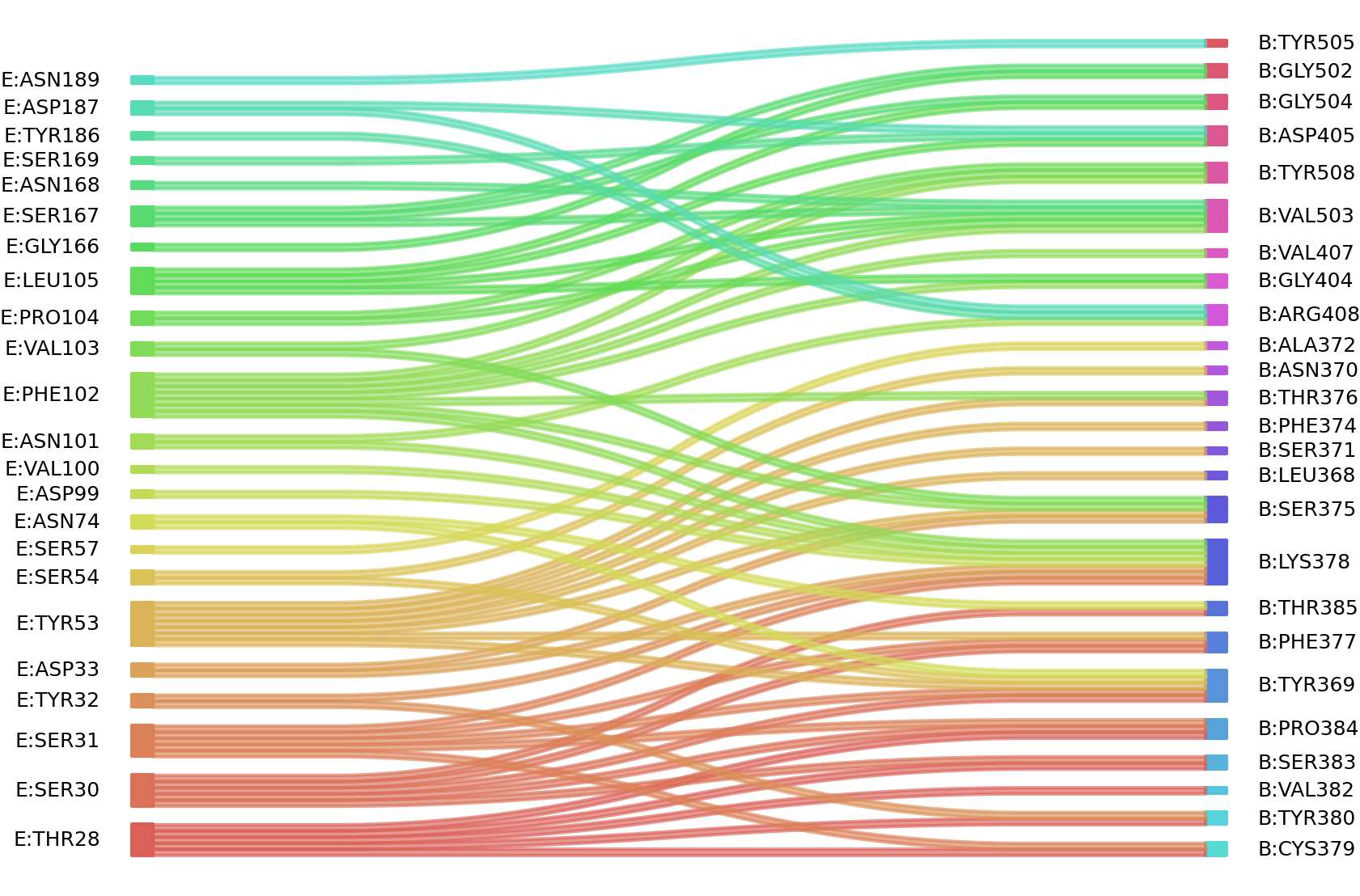
2、We defined interacting paratope-epitope residues by a distance cutoff of < 6 Å . Two amino acids are considered as interacting residues if they have at least one atom within a distance of 6 Å from any atom.
Interacting residues (Atom distance based)