Details
Structure visualisation
Entry information
| Complex | |
| AACDB_ID: | 438 |
| PDBID: | 3BT2 |
| Chains: | HL_U |
| Organism: | Homo sapiens, Mus musculus |
| Method: | XRD |
| Resolution (Å): | 2.50 |
| Reference: | 10.1038/nsmb.1404 |
| Antibody | |
| Antibody: | 3BT2 Fab |
| Antibody mutation: | No |
| INN (Clinical Trial): | |
| Antigen | |
| Antigen: | Urokinase plasminogen activator surface receptor |
| Antigen mutation: | No |
| Durg Target: | Q03405 |
Sequence information
Antibody
Heavy Chain: H
Mutation: NULL
| >3BT2_H|Chain D[auth H]|anti-uPAR antibody, heavy chain|Mus musculus (10090) GVKLQQSGPEVVKPGASVKISCKASGYSFTNFYIHWVKQRPGQGLEWIGWIFHGSDNTEYNEKFKDKATLTADTSSSTAYMQLSSLTSEDSAVYFCARWGPHWYFDVWGQGTTVTVSSAKTTPPSVYPLAPGNSMVTLGCLVKGYFPEPVTVTWNSGSLSSGVHTFPAVLQSDLYTLSSSVTVPSSTWPSETVTCNVAHPASSTKVDKKIAAAG |
Light Chain: L
Mutation: NULL
| >3BT2_L|Chain C[auth L]|anti-uPAR antibody, light chain|Mus musculus (10090) DIVLTQSPDITAASLGQKVTITCSASSSVSYMHWYQQKSGTSPKPWIFEISKLASGVPARFSGSGSGTSYSLTISSMEAEDAAIYYCQQWNYPFTFGGGTKLEIKRADAAPTVSIFPPSSEQLTSGGASVVCFLNNFYPKDINVKWKIDGSERQNGVLNSWTDQDSKDSTYSMSSTLTLTKDEYERHNSYTCEATHKTSTSPIVKSFNRNEC |
Antigen
Chain: U
Mutation: NULL
| >3BT2_U|Chain E[auth U]|Urokinase plasminogen activator surface receptor|Homo sapiens (9606) RSLRCMQCKTNGDCRVEECALGQDLCRTTIVRLWEEGEELELVEKSCTHSEKTNRTLSYRTGLKITSLTEVVCGLDLCNQGNSGRAVTYSRSRYLECISCGSSDMSCERGRHQSLQCRSPEEQCLDVVTHWIQEGEEGRPKDDRHLRGCGYLPGCPGSNGFHNNDTFHFLKCCNTTKCNEGPILELENLPQNGRQCYSCKGNSTHGCSSEETFLIDCRGPMNQCLVATGTHEPKNQSYMVRGCATASMCQHAHLGDAFSMNHIDVSCCTKSGCNHPDLDVQYR |
Interaction
1、Solvent accessible surface areas (SASA) were calculated (Naccess V2.1.1) for each residue in antibody and antigen, respectively. The residues with SASA loss in binding of more than 1Å2 were classified as interacting residues.
Interacting residues (ΔSASA based)
| Chain residues position delta_SASA
: residuesposition L: SER31 TYR32 TRP91 ASN92 TYR93 PHE96 H: TYR33 TRP50 HIS52 ASP55 ASN56 THR57 GLU58 TRP95 HIS98 TRP99 U: PRO180 GLU185 ASN186 LEU187 PRO188 GLN189 ASN190 GLY191 ARG192 ARG216 GLY217 PRO218 ASN220 GLN221 ALA244 THR267 LYS268 SER269 |
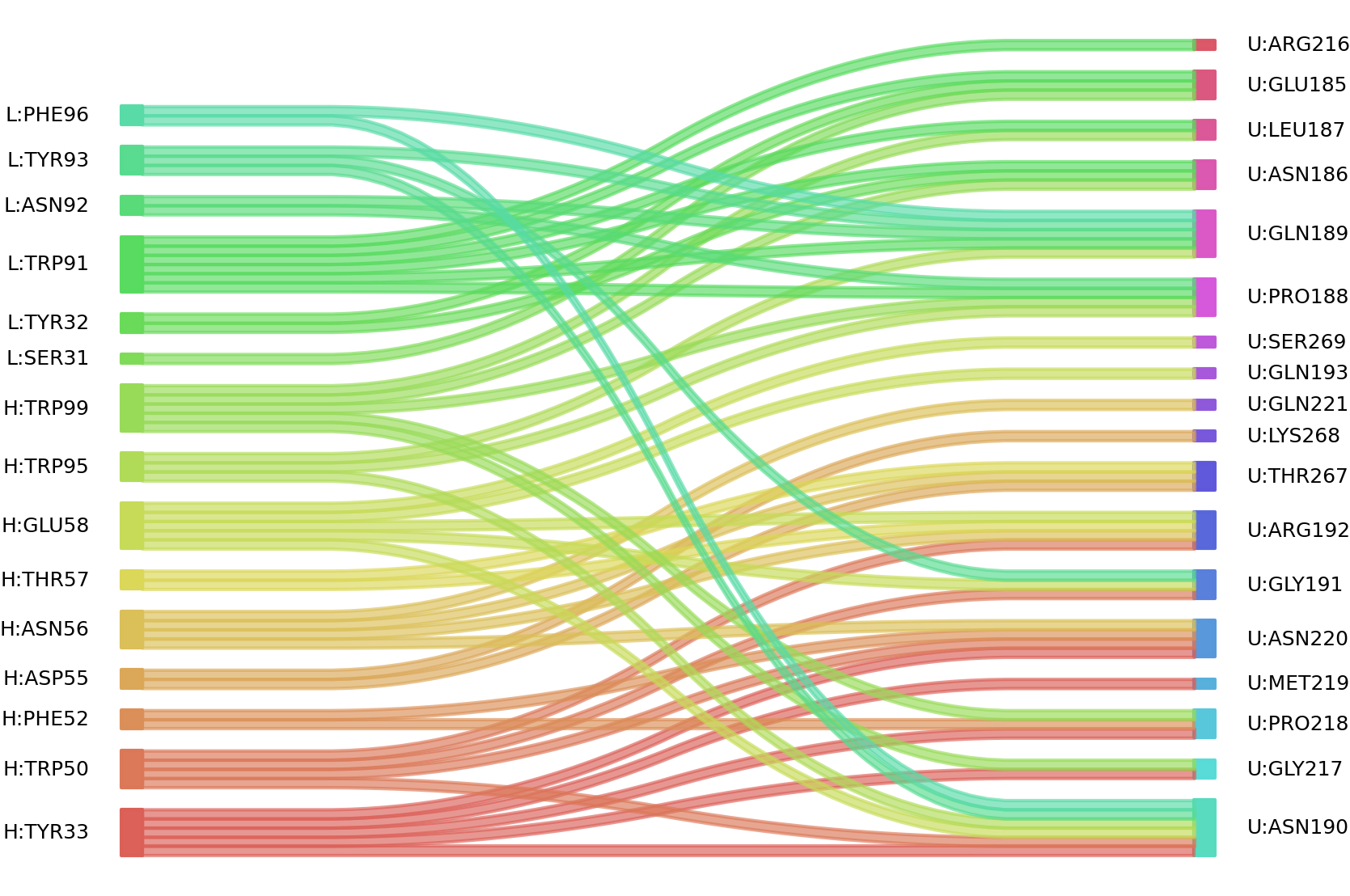
2、We defined interacting paratope-epitope residues by a distance cutoff of < 6 Å . Two amino acids are considered as interacting residues if they have at least one atom within a distance of 6 Å from any atom.
Interacting residues (Atom distance based)