Details
Structure visualisation
Entry information
| Complex | |
| AACDB_ID: | 4166 |
| PDBID: | 7LOK |
| Chains: | ML_C |
| Organism: | Human immunodeficiency virus 1, Homo sapiens, Leiurus quinquestriatus hebraeus |
| Method: | EM |
| Resolution (Å): | 3.90 |
| Reference: | 10.1038/s41467-021-21816-x |
| Antibody | |
| Antibody: | 17b Fab |
| Antibody mutation: | No |
| INN (Clinical Trial): | |
| Antigen | |
| Antigen: | HIV-1 BG505 SOSIP.664 gp120 |
| Antigen mutation: | No |
| Durg Target: | |
Sequence information
Antibody
Heavy Chain: M
Mutation: NULL
| >7LOK_M|Chain F[auth K], H[auth M]|17b Fab Heavy Chain|Homo sapiens (9606) EVQLVESGAEVKKPGSSVKVSCKASGDTFIRYSFTWVRQAPGQGLEWMGRIITILDVAHYAPHLQGRVTITADKSTSTVYLELRNLRSDDTAVYFCAGVYEGEADEGEYDNNGFLKHWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEPKSCDKHHHHHH |
Light Chain: L
Mutation: NULL
| >7LOK_L|Chain E[auth J], G[auth L]|17b Fab Light Chain|Homo sapiens (9606) DIVMTQSPATLSVSPGERATLSCRASESVSSDLAWYQQKPGQAPRLLIYGASTRATGVPARFSGSGSGAEFTLTISSLQSEDFAVYYCQQYNNWPPRYTFGQGTRLEIKRTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRG |
Antigen
Chain: C
Mutation: NULL
| >7LOK_C|Chain A, C, L[auth E]|Envelope glycoprotein BG505 SOSIP.664 gp120|Human immunodeficiency virus 1 (11676) NLWVTVYYGVPVWKDAETTLFCASDAKAYETEKHNVWATHACVPTDPNPQEIHLENVTEEFNMWKNNMVEQMHTDIISLWDQSLKPCVKLTPLCVTLQCTNVTNNITDDMRGELKNCSFNMTTELRDKKQKVYSLFYRLDVVQINENQGNRSNNSNKEYRLINCNTSAITQACPKVSFEPIPIHYCAPAGFAILKCKDKKFNGTGPCPSVSTVQCTHGIKPVVSTQLLLNGSLAEEEVMIRSENITNNAKNILVQFNTPVQINCTRPNNNTRKSIRIGPGQAFYATGDIIGDIRQAHCNVSKATWNETLGKVVKQLRKHFGNNTIIRFANSSGGDLEVTTHSFNCGGEFFYCNTSGLFNSTWISNTSVQGSNSTGSNDSITLPCRIKQIINMWQRIGQAMYAPPIQGVIRCVSNITGLILTRDGGSTNSTTETFRPGGGDMRDNWRSELYKYKVVKIEPLGVAPTRCKRRVVGRRRRRR |
Interaction
1、Solvent accessible surface areas (SASA) were calculated (Naccess V2.1.1) for each residue in antibody and antigen, respectively. The residues with SASA loss in binding of more than 1Å2 were classified as interacting residues.
Interacting residues (ΔSASA based)
| Chain residues position delta_SASA
: residuesposition L: TRP94 PRO95 M: ILE52 ILE54 LEU55 VAL57 HIS59 ASP105 GLU106 GLY107 GLU108 TYR109 ASP110 ASN111 C: CYS119 VAL120 LEU122 ALA200 THR202 GLN203 ALA204 CYS205 ARG327 LEU369 ILE420 LYS421 GLN422 ILE423 GLN432 ALA433 MET434 ALA436 PRO437 |
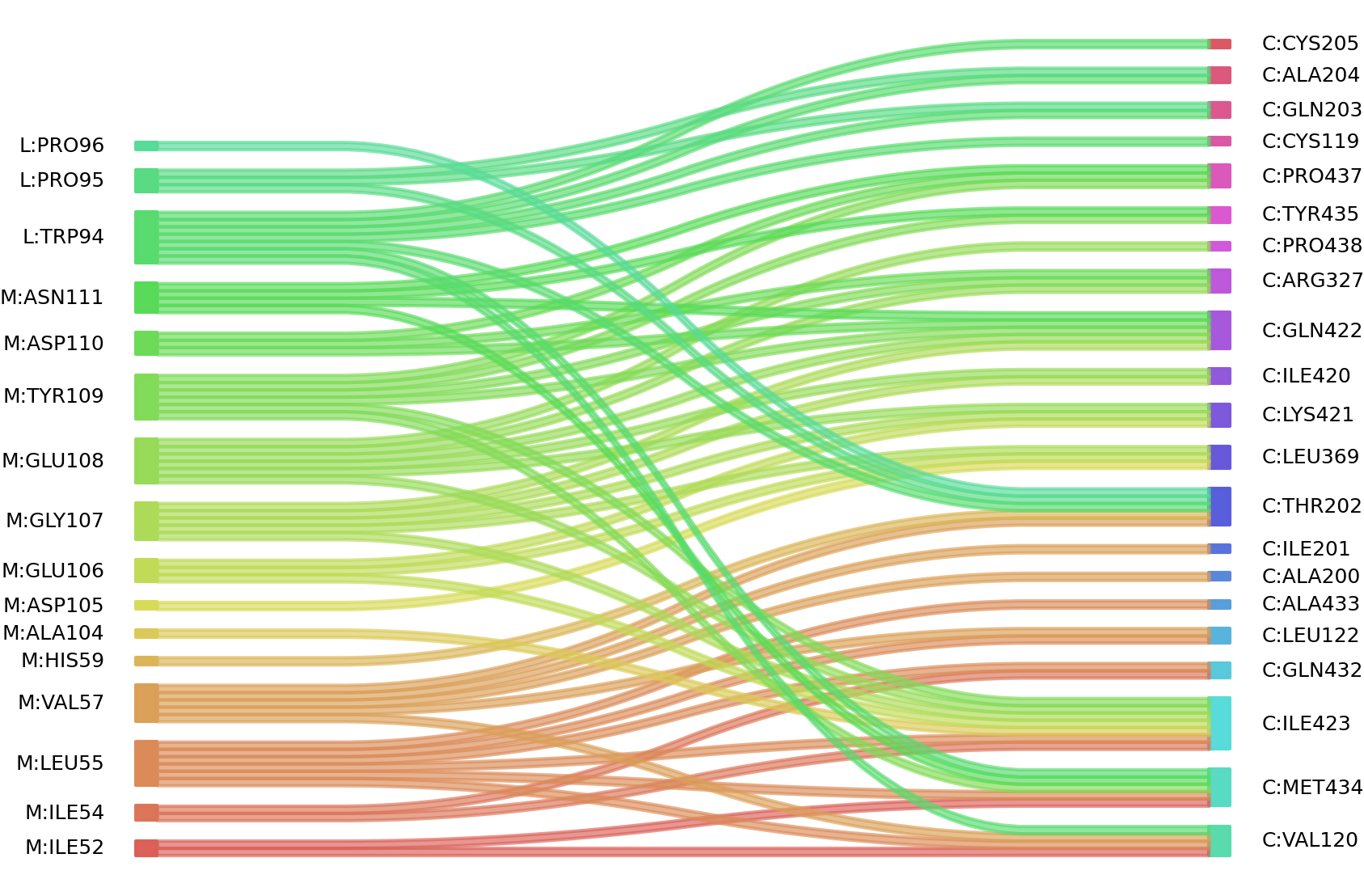
2、We defined interacting paratope-epitope residues by a distance cutoff of < 6 Å . Two amino acids are considered as interacting residues if they have at least one atom within a distance of 6 Å from any atom.
Interacting residues (Atom distance based)