Details
Structure visualisation
Entry information
| Complex | |
| AACDB_ID: | 2496 |
| PDBID: | 6MW9 |
| Chains: | KL_J |
| Organism: | Eastern equine encephalitis virus, Mus musculus |
| Method: | EM |
| Resolution (Å): | 7.30 |
| Reference: | 10.1016/j.celrep.2018.11.067 |
| Antibody | |
| Antibody: | EEEV-3 Fab |
| Antibody mutation: | No |
| INN (Clinical Trial): | |
| Antigen | |
| Antigen: | chimeric Eastern Equine Encephalitis Virus E2 |
| Antigen mutation: | No |
| Durg Target: | |
Sequence information
Antibody
Heavy Chain: K
Mutation: NULL
| >6MW9_K|Chain C, G, K, O|EEEV-3 antibody heavy chain|Mus musculus (10090) QIQLVQSGREVKNPGETVKISCKASGYTFTEYPMLWVKQAPGKGFRWMGLIYTNTGEPTYAEEFKGRFVFSLEISASTAYLQINNLTNEDTATYFCVRDYFISLDYWGQGTTLTVSSAKTTAPSVYPLAPVCGGTTGSSVTLGCLVKGYFPEPVTLTWNSGSLSSGVHTFPALLQSGLYTLSSSVTVTSNTWPSQTITCNVAHPASSTKVDKKIESRR |
Light Chain: L
Mutation: NULL
| >6MW9_L|Chain D, H, L, P|EEEV-3 antibody light chain|Mus musculus (10090) QAVVTQESALTTSPGETVTLTCRSNIGAVTSSNCANWVQEKPDHFFTGLIGDTNNRRSGVPARFSGSLIGDKAALTITGAQTEDEAIYFCALWYNNLWVFGGGTKLTVLGQPKSSPSVTLFPPSSEELETNKATLVCTITDFYPGVVTVDWKVDGTPVTQGMETTQPSKQSNNKYMASSYLTLTARAWERHSSYSCQVTHEGHTVEKSLSRADC |
Antigen
Chain: J
Mutation: NULL
| >6MW9_J|Chain B, F, J, N|E2|Eastern equine encephalitis virus (11021) DLDTHFTQYKLARPYIADCPNCGHSRCDSPIAIEEVRGDAHAGVIRIQTSAMFGLKTDGVDLAYMSFMNGKTQKSIKIDNLHVRTSAPCSLVSHHGYYILAQCPPGDTVTVGFHDGPNRHTCTVAHKVEFRPVGREKYRHPPEHGVELPCNRYTHKRADQGHYVEMHQPGLVADHSLLSIHSAKVKITVPSGAQVKYYCKCPDVREGITSSDHTTTCTDVKQCRAYLIDNKKWVYNSGRLPRGEGDTFKGKLHVPFVPVKAKCIATLAPEPLVEHKHRTLILHLHPDHPTLLTTRSLGSDANPTRQWIERPTTVNFTVTGEGLEYTWGNHPPKRVWAQESGEGNPHGWPHEVVVYYYNRYPLTTIIGLCTCVAIIMVSCVTSVWLLCRTRNLCITPYKLAPNAQVPILLALLCCIKPTRA |
Interaction
1、Solvent accessible surface areas (SASA) were calculated (Naccess V2.1.1) for each residue in antibody and antigen, respectively. The residues with SASA loss in binding of more than 1Å2 were classified as interacting residues.
Interacting residues (ΔSASA based)
| Chain residues position delta_SASA
: residuesposition K: PRO4033 TRP4047 LEU4050 TYR4052 ASN4054 THR4055 GLU4057 THR4059 ASP4099 ILE4102 L: SER4531 SER4532 CYS4534 TRP4593 ASN4595 ASN4596 TRP4598 J: HIS2662 SER2663 LEU2665 SER2666 ILE2667 HIS2668 SER2669 ALA2670 LYS2671 LYS2673 THR2675 HIS2700 THR2702 THR2705 ASP2706 VAL2707 LYS2708 |
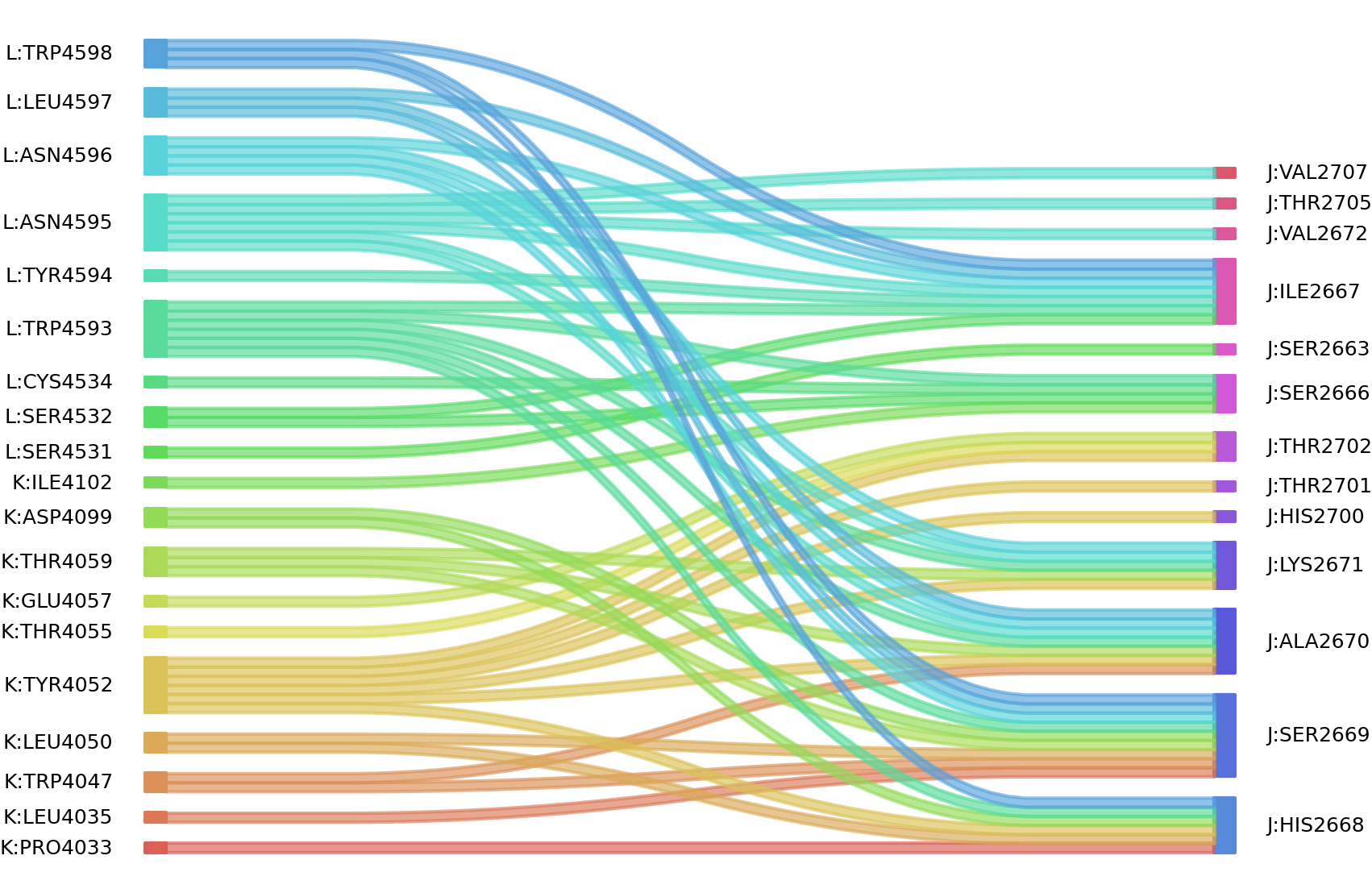
2、We defined interacting paratope-epitope residues by a distance cutoff of < 6 Å . Two amino acids are considered as interacting residues if they have at least one atom within a distance of 6 Å from any atom.
Interacting residues (Atom distance based)