Details
Structure visualisation
Entry information
| Complex | |
| AACDB_ID: | 1107 |
| PDBID: | 4YDJ |
| Chains: | HL_G |
| Organism: | Human immunodeficiency virus 1, Homo sapiens |
| Method: | XRD |
| Resolution (Å): | 2.31 |
| Reference: | 10.1016/j.cell.2015.05.007 |
| Antibody | |
| Antibody: | 44-VRC13.01 Fab |
| Antibody mutation: | No |
| INN (Clinical Trial): | |
| Antigen | |
| Antigen: | HIV-1 clade AE strain 93TH057 gp120 |
| Antigen mutation: | No |
| Durg Target: | |
Sequence information
Antibody
Heavy Chain: H
Mutation: NULL
| >4YDJ_H|Chain B[auth A], E[auth H]|HEAVY CHAIN OF ANTIBODY 44-VRC13.01|Homo sapiens (9606) QVQLVQPGTAMKSLGSSLTITCRVSGDDLGSFHFGTYFMIWVRQAPGQGLEYMGGILPSTKTPTYAHKFRGRVSISAPGVPPVLSLALTNLTYDDTATYFCARERGRHFEPKNRDNLEGKFFDLWGRGTFVRVSPASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSC |
Light Chain: L
Mutation: NULL
| >4YDJ_L|Chain C[auth B], F[auth L]|LIGHT CHAIN OF ANTIBODY 44-VRC13.01|Homo sapiens (9606) QSALTQPASVSGSPGQSINISCAGRSDRVSWYQQRPNGVPKLLMFDVYRRPSGVSDRFSGSHSGDTAFLTISGLQTEDEADYYCTSHPYAFGAGTKVNVLRQPKANPTVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHKSYSCQVTHEGSTVEKTVAPTECS |
Antigen
Chain: G
Mutation: NULL
| >4YDJ_G|Chain A[auth G], D[auth I]|Envelope glycoprotein gp160,Envelope glycoprotein gp160|Human immunodeficiency virus 1 (11676) VWKDADTTLFCASDAKAHETEVHNVWATHACVPTDPNPQEIHLENVTENFNMWKNNMVEQMQEDVISLWDQSLQPCVKLTGGSVIKQACPKISFDPIPIHYCTPAGYVILKCNDKNFNGTGPCKNVSSVQCTHGIKPVVSTQLLLNGSLAEEEIIIRSENLTNNAKTIIVHLNKSVEINCTRPSNGGSGSGGDIRKAYCEINGTKWNKVLKQVTEKLKEHFNNKTIIFQPPSGGDLEITMHHFNCRGEFFYCNTTQLFNNTCIGNETMKGCNGTITLPCKIKQIINMWQGTGQAMYAPPIDGKINCVSNITGILLTRDGGANNTSNETFRPGGGNIKDNWRSELYKYKVVQIE |
Interaction
1、Solvent accessible surface areas (SASA) were calculated (Naccess V2.1.1) for each residue in antibody and antigen, respectively. The residues with SASA loss in binding of more than 1Å2 were classified as interacting residues.
Interacting residues (ΔSASA based)
| Chain residues position delta_SASA
: residuesposition H: LEU29 GLY30 SER31 HIS33 PHE34 VAL74 GLU95 ARG96 GLY97 ARG98 HIS99 L: ASP31 ARG32 ARG53 G: TRP96 LYS97 GLU275 ASN279 ASN280 ALA281 LYS282 THR283 ARG327 PRO364 SER365 GLY366 GLY367 ASP368 LEU369 ILE371 THR372 MET373 TYR384 LYS419 ILE420 LYS421 ILE423 THR455 GLY458 GLY459 ARG469 PRO470 GLY472 GLY473 ASN474 ASP477 ARG480 |
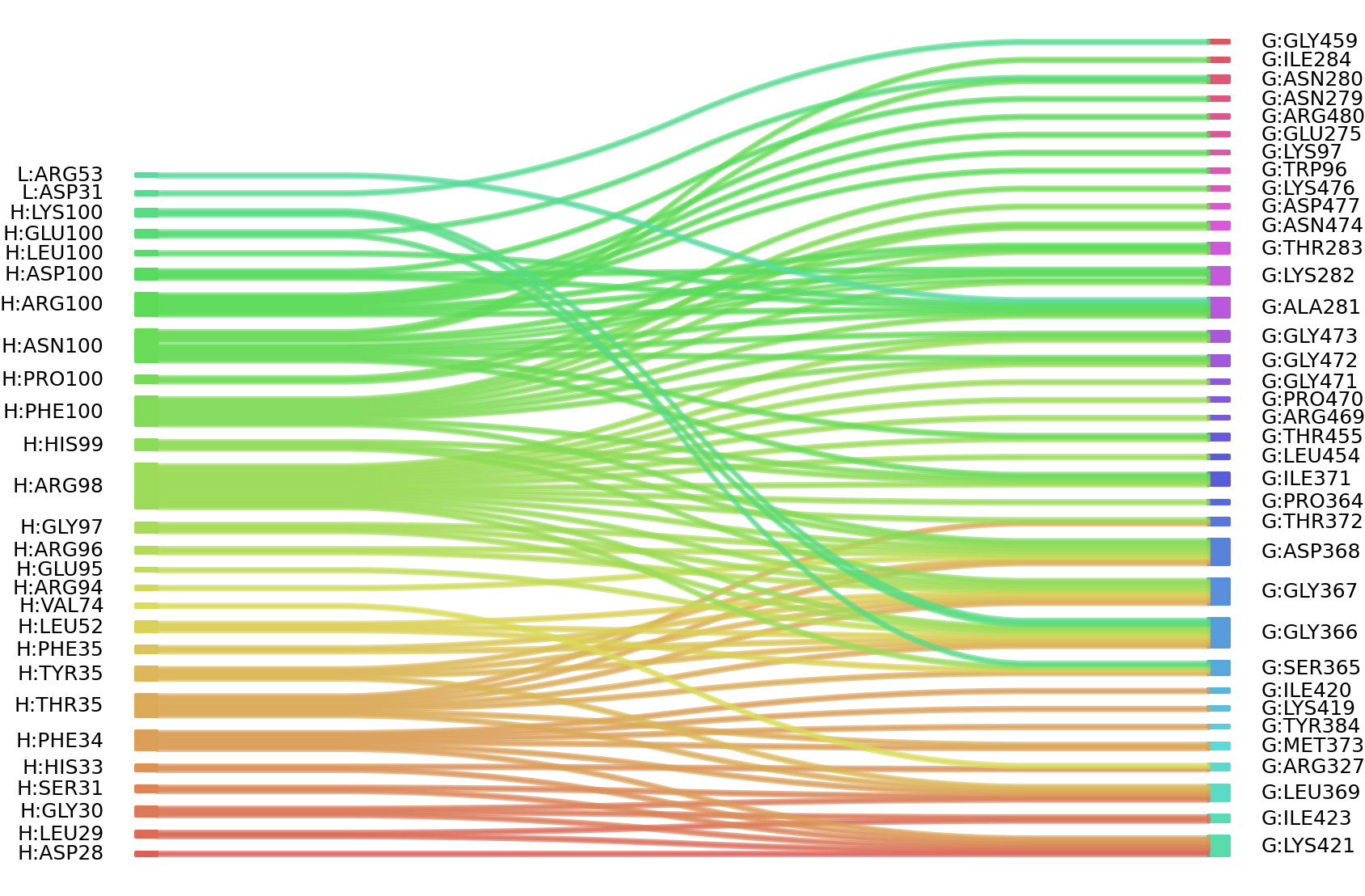
2、We defined interacting paratope-epitope residues by a distance cutoff of < 6 Å . Two amino acids are considered as interacting residues if they have at least one atom within a distance of 6 Å from any atom.
Interacting residues (Atom distance based)